Given strings "ABAB" and "BABA"...
A dynamic programming style algorithm constructs a matrix of the substring lengths.
| | | A | B | A | B |
| | O | O | O | O | O |
| B | O | O | 1 | O | 1 |
| A | O | 1 | 0 | 2 | 0 |
| B | O | 0 | 2 | 0 | 3 |
| A | O | 1 | 0 | 3 | 0 |
In this matrix, the 5th column tests the substring ABA of the ABAB sequence. So cell [5, 5] compares ABA to BAB ... the lcs is BA, length 2.
The other approach is to construct a generalized suffix tree for the strings, then find the deepest internal nodes which have leaf nodes from all the strings in the subtree below it.
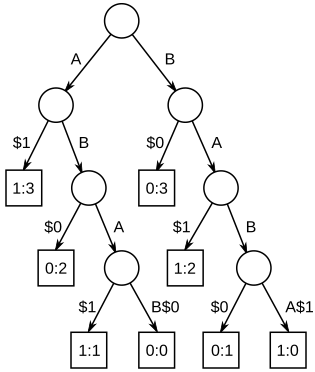
A Simple Suffix Tree Implementation:
Dijkstras Algorithm for shortest path:
Or, here's a more complete implementation.